Image Processing - Method 1
Protocol
Image Capture
Histology sections were captured as digital images as described in Image Capture - Configuration A
3D Reconstruction - reconstruct
The processing and sectioning of the embryo, which includes chemical and mechanical insult to the tissues which causes systematic and random distortions of each section. We wish to recover a representative model of the embryo which can be re-sectioned digitally and produce a realistic image of a section. For this we need to correct all the minor random distortions so that the tissues re-align accurately. We acheive this by warping (see below). The warping process is slow and cpu intensive and requires that the adjacent sections are already close in alignment so that all distortions are small. We therefore apply a rough alignment during digitisation followed by manual/automatic registration. The program developed for registration is reconstruct which also provides a convenient GUI to organise the section files, review the images, normalise the grey-values and re-scale if required. This program is also being developed as a means of reconstructing voxel images with multiple images per section as well as incorporating 2D tie-point warping based on thin-plate splines.
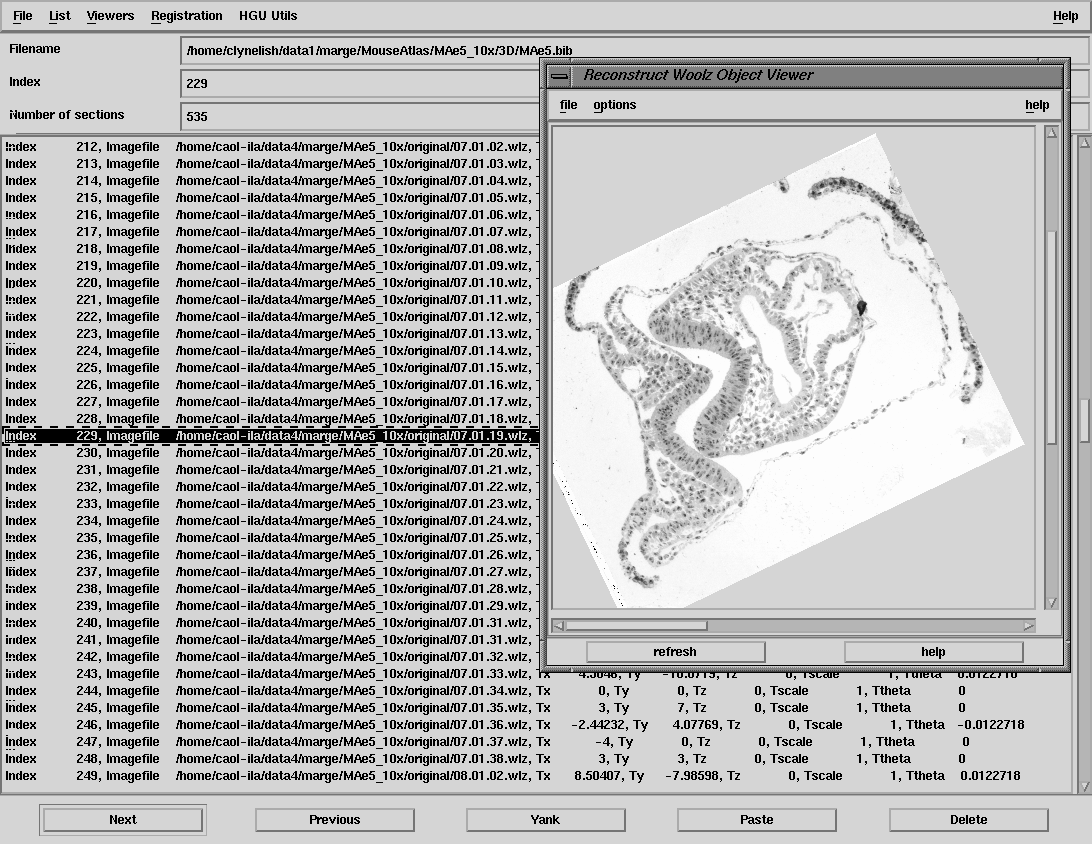
The underlying philosophy adopted for the reconstruct program is that the source data files are left unchanged and all modifications are defined within a text file so that the 3D voxel image can be created as required from the originals. The text file is a series of records which define the relative position and transformations for each section and is modelled in the bibtex style and is therefore referred to as the "bibfile". This style of file is also used as the flatfile format for the full anatomy and gene-expression database. Reconstruct can be used to create the bibfile from a selected list of image files and will then provide an interface to allow editing of the records. All the usual operations of cut, copy and paste are provided as well as editing of the record itself, either by direct interative manipulation of the images or by editing the individual values of translation, rotation, scale etc of each section.
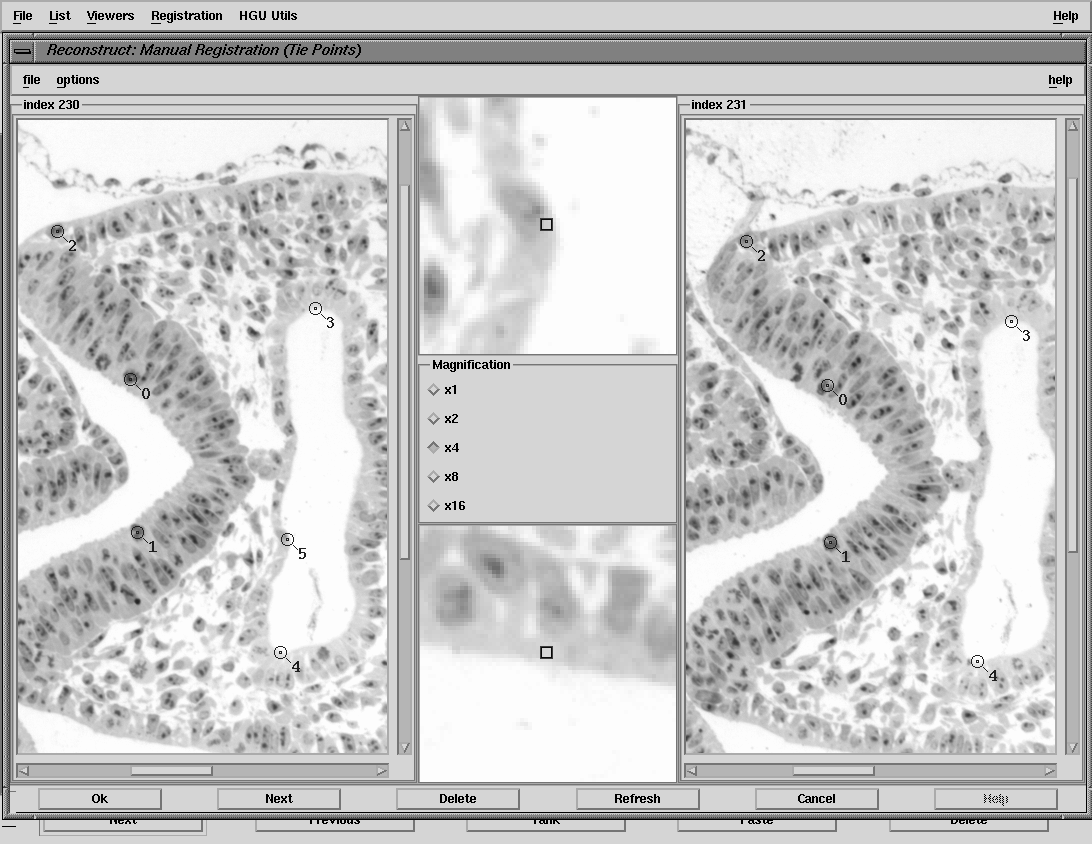
Reconstruct allows convenient browsing through digitised images and is therefore used for the initial task of reviewing the section images to check for accidental repeats, out-of-focus sections, damaged or torn sections. This review process may invoke re-digitisation of some sections and is used to generate the final list of images to be used to create the voxel model. Where sections are missing or unusable for some reason then adjacent sections are substituted and warped to be "halfway" between the sections above and below. This warping is done using the thin-plate spline interpolation function from a series of tie-points. This functionality is built into reconstruct (currently in prototype mode).
Output operations from reconstruct allow the creation of the bibfile, building the voxel image with options on volume of interest and normalisation of the grey-values. Grey-value normalisation is important because the sections (especially plastic embedded sections) do not always have consistent stain penetration and possibly variable section thickness. This means that some sections appear darker than others and this creates artifacts in sections through the data not parallel to the original sectioning planes. Normalisation is an automatic process which matches the grey-level histograms of each plane and then resets the grey-values to match a mean distribution.
| reconstruct interface - reviewing the sections and showing the current transformations |
 |
| reconstruct interface - tie point interface |
 |
Distortion Correction - warpstack
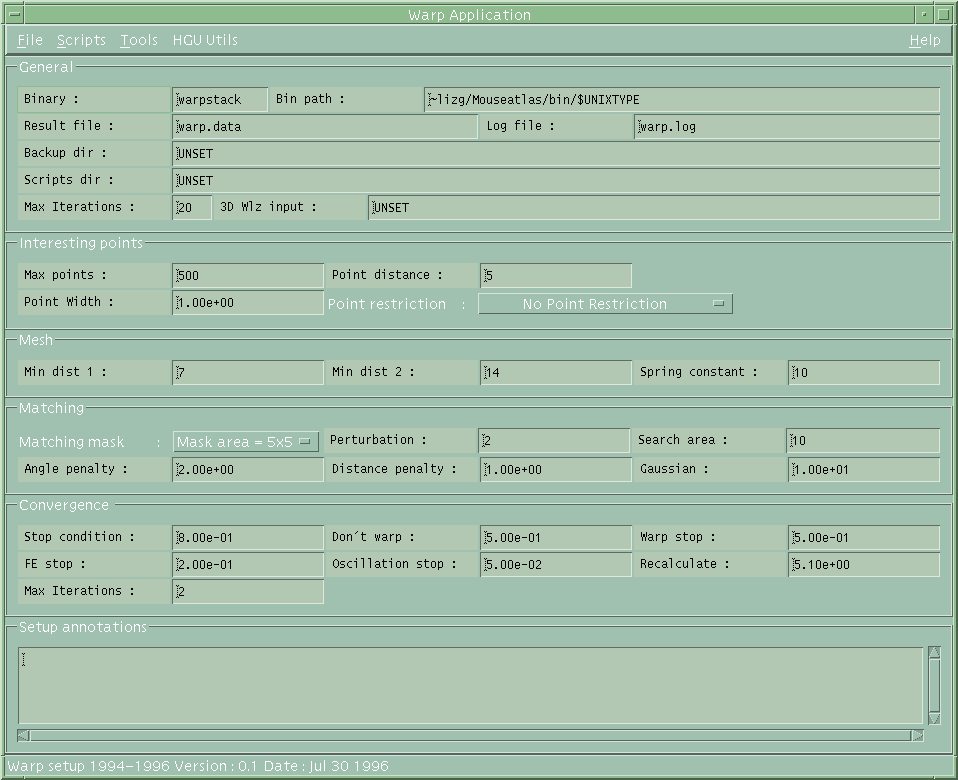
Once the section images have been reconstructed to an aligned and normalised voxel image, this image is passed to warpstack to correct the sectioning distortions and to refine the automated alignment from reconstruct. The basic idea is to smooth the biological structures visible in the images with the underlying assumption that the section sampling size (2 or 7 microns) is small with respect to the typical radii of curvature of the surfaces of biological structures. This is not always true but because of the elastic model employed and the matching redundancy in the method, occasional errors do not significantly affect the final result.
The basic idea is to model each section as a thin elastic sheet with its own internal elastic properties. In most cases we assume uniform elasticity but the methods allows for any arbitrary non-linear elaticity functions which can vary for different structures eg neural tissue, membranes, mesenchyne etc. For each section a number of interesting points (400-500) are defined, scattered over the image and associated with points that have a useful matchable feature which in this case is defined to be points of high gradient in grey-value, or grey-value variance. These points are then matched both to the section above and the section below to find both a position of best match and an associated confidence. The position of best match allows a "spring" to be attached between the images and the confidence determines the strength of the spring. The section image is then deformed until the forces generated by the springs and the internal strains are in equilibrium. This equilibrium is found using the Finite Element Method (FEM). This is repeated for each section (except the top and bottom) then the whole calculation repeated iteratively until the calculation converges.
There are a number of points which need consideration during the warping process. One is that most interesting points lie on "edges" in the image and therefore there is an "aperture problem" in matching, ie the match is in general badly constrained parallel to the edge direction. For this reason the match is implemented as only constrained perpendicular to the edge and otherwise allowed to "slide along". The assumption is that other points matched in the image will force convergence in the parallel direction. The other point is that like all automatic matching algorithms dealing with real imge data, some match errors occur. In most cases this is rendered harmless by the calculation of a match confidence which means the spring strength is negligible and therefore the tissue is not affected. In other cases it is the elastic properties of the tissue which prevent undue distortion by an erroneous match, because the mismatch will be conteracted by the surrounding correct matches. This is why a high redundancy in the number of match points is important.
The whole process is fully automatic once the parameters have been set and for large data sets can take many days cpu time on a Sun Microsystems Sparc-10 workstation. For this reason we have also developed parallel version of the program which can run on large scale parallel machine (eg the T3D in Edinburgh) or in distributed fashion. The parallelism is achieved using MPI and because of the easy way the problem can be split into large cpu intensive chunks with little communication, near linear speedup is possible.
| warpstack control interface |
 |