MGEIR - The Mouse Gene Expression Information Resource
EMAGE is part of the Mouse Gene Expression Information Resource (MGEIR) project which is a collaboration between EMAP and the Gene Expression Database (GXD) project at Mouse Genome Informatics (MGI) at the Jackson Laboratory, USA.
The ultimate aim of the MGEIR is to provide a unified resource that combines text-based and spatial-based methods to store, display, and analyse mouse developmental gene expression information.
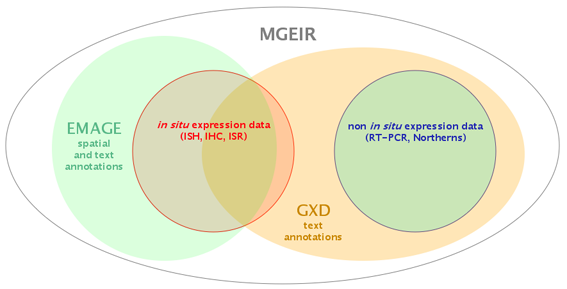
Both GXD and EMAGE obtain mouse embryo gene expression data from the literature and by direct data submission from external labs/consortia. One important difference between GXD and EMAGE is that whereas EMAGE only includes expression data from in situ techniques such as ISH and IHC, the GXD also incorporates data from 'non-spatial' expression profiling techniques such as RT-PCRs, Northern blots etc.
Both GXD and EMAGE convert the unstructured information received from each external source into a standardised description that is available for database storage and query. During this process we adhere to common guidelines that result in consistent descriptions that can be shared between the two databases.
The critical difference between the methods used by GXD and EMAGE to describe patterns of gene expression, is that whereas both use a standardised text description against the EMAP anatomy ontology, EMAGE also performs a spatial annotation.
The creation of MGEIR will provide integration of the data annotated at GXD and EMAGE into one common queryable resource.

 icon to keep this page displayed.)
icon to keep this page displayed.)


