|
|
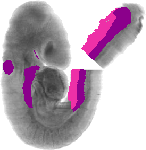
Gata3 |
|
 |
 |
|
TS14 |
9.5-10.0 dpc |
EMAGE:573
view entry |
ISH |
 |
|
|
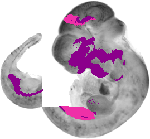
Gata3 |
|
 |
 |
|
TS18 |
10.5 dpc |
EMAGE:2808
view entry |
ISH |
 |
|
|
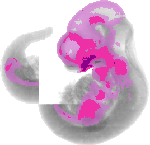
Gata3 |
|
 |
 |
|
TS17 |
10.5 dpc |
EMAGE:2809
view entry |
ISH |
 |
|
|
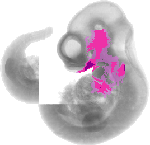
Gata3 |
|
 |
 |
|
TS17 |
10.5 dpc |
EMAGE:3312
view entry |
ISH |
 |
|
|
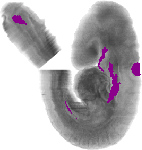
Gata3 |
|
 |
 |
- |
TS14 |
9.0 dpc |
EMAGE:3921
view entry |
ISH |
 |
|
|
Gata3 |
|
 |
 |
|
TS18 |
10.5 dpc |
EMAGE:3922
view entry |
ISH |
 |
|
|
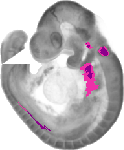
Gata3 |
|
 |
 |
|
TS15 |
9.5 dpc |
EMAGE:3923
view entry |
ISH |
 |
|
|
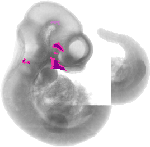
Gata3 |
|
 |
 |
|
TS17 |
10.5 dpc |
EMAGE:4575
view entry |
ISH |
 |
|
|
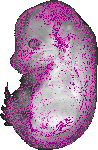
Gata3 |
|
 |
 |
- |
TS23 |
14.5 dpc |
EMAGE:9422
view entry |
ISH |
 |
|
|
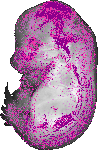
Gata3 |
|
 |
 |
| Go to annotation details |
 |
 |
bladder |
 |
adrenal medulla |
 |
spinal cord mantle layer |
|
 |
perioptic mesenchyme |
 |
nasal cavity |
 |
cervical ganglion |
 |
diencephalon lateral wall mantle layer |
 |
midbrain mantle layer |
 |
pons mantle layer |
 |
kidney calyx |
 |
cervico-thoracic ganglion |
 |
inner ear |
 |
larynx |
 |
medulla oblongata basal plate mantle layer |
 |
skin |
 |
pharyngo-tympanic tube |
 |
urinary system |
 |
thymus primordium |
 |
vibrissa |
 |
urethra of male |
 |
thoracic ganglion |
|
|
TS23 |
14.5 dpc |
EMAGE:11708
view entry |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)