|
|
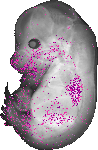
Lama5 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
optic stalk |
 |
vibrissa |
 |
lung |
|
 |
kidney calyx |
 |
stomach |
 |
lateral ventricle choroid plexus |
 |
cochlea |
 |
upper jaw incisor |
 |
kidney pelvis |
 |
esophagus |
 |
saccule |
 |
sublingual gland primordium |
 |
pharyngo-tympanic tube |
 |
pituitary gland |
 |
lower jaw incisor |
 |
male reproductive system |
 |
oral epithelium |
 |
cornea epithelium |
 |
urinary system |
 |
larynx |
 |
lower jaw molar |
 |
midgut |
 |
urethra of male |
 |
bladder |
 |
hindgut |
 |
upper jaw molar |
 |
trachea |
 |
submandibular gland primordium |
 |
epidermis |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
rectum |
 |
naris |
 |
nasal cavity olfactory epithelium |
 |
thyroid gland |
|
|
EMAGE:11991
view entry |
wild-type |
- |
ISH |
 |
|
|
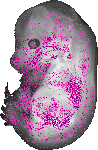
Itga3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.159 |
- |
EMAGE:21631
view entry |
wild-type |
- |
ISH |
 |
|
|
Bambi |
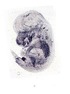 |
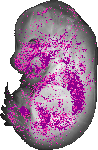 |
 |
TS23 |
14.5 dpc |
0.157 |
- |
EMAGE:9050
view entry |
wild-type |
- |
ISH |
 |
|
|
Krt19 |
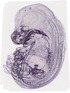 |
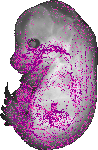 |
 |
TS23 |
14.5 dpc |
0.152 |
| Go to annotation details |
 |
 |
lower jaw molar |
 |
midgut |
 |
nasal cavity epithelium |
|
 |
pharyngo-tympanic tube |
 |
vibrissa |
 |
pharynx |
 |
trachea |
 |
bladder |
 |
kidney calyx |
 |
urethra of male |
 |
submandibular gland primordium |
 |
right lung |
 |
thymus primordium |
 |
pleural cavity |
 |
skin |
 |
peritoneal component |
 |
inner ear |
 |
stomach |
 |
larynx |
 |
left lung |
 |
oral epithelium |
 |
upper jaw molar |
 |
naso-lacrimal duct |
 |
upper jaw incisor |
 |
eyelid |
 |
urogenital mesentery |
 |
pleura |
 |
rectum |
 |
esophagus |
 |
pericardium |
 |
ureter |
 |
nasal cavity olfactory epithelium |
 |
lower jaw incisor |
 |
anterior naris |
 |
external naris |
|
|
EMAGE:6677
view entry |
wild-type |
- |
ISH |
 |
|
|
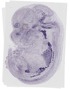
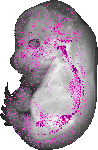
Maoa |
 |
 |
 |
TS23 |
14.5 dpc |
0.149 |
| Go to annotation details |
 |
 |
metencephalon |
 |
olfactory lobe |
 |
trigeminal V ganglion |
|
 |
adrenal cortex |
 |
vestibulocochlear VIII ganglion |
 |
vagus X ganglion |
 |
cervical ganglion |
 |
medulla oblongata |
 |
lung |
 |
midbrain ventricular layer |
 |
vibrissa |
 |
thoracic ganglion |
 |
dorsal root ganglion |
 |
pharyngo-tympanic tube |
 |
midbrain mantle layer |
 |
cervico-thoracic ganglion |
 |
cochlea |
|
|
EMAGE:10938
view entry |
wild-type |
- |
ISH |
 |
|
|
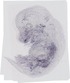
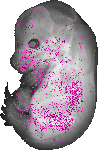
Itga8 |
 |
 |
 |
TS23 |
14.5 dpc |
0.149 |
| Go to annotation details |
 |
 |
midgut |
 |
lateral ventricle choroid plexus |
 |
metencephalon part of 4th ventricle choroid plexus |
|
 |
nasal septum |
 |
peritoneal component |
 |
pericardial cavity |
 |
pleural cavity |
 |
diaphragm |
 |
upper lip |
 |
lower lip |
|
|
EMAGE:13973
view entry |
wild-type |
- |
ISH |
 |
|
|
Frk |
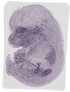 |
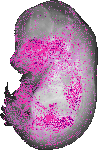 |
 |
TS23 |
14.5 dpc |
0.148 |
| Go to annotation details |
 |
 |
midgut |
 |
urethra of male |
 |
mandible |
|
 |
rectum |
 |
limb |
 |
axial skeleton |
 |
stomach |
 |
palatal shelf |
 |
naris |
 |
nasal cavity respiratory epithelium |
 |
vibrissa |
 |
lung |
 |
kidney pelvis |
 |
trachea |
 |
bladder |
 |
cornea |
 |
adenohypophysis |
 |
kidney calyx |
 |
tongue |
 |
pharyngo-tympanic tube |
 |
nasal cavity olfactory epithelium |
 |
hindlimb |
|
|
EMAGE:11087
view entry |
wild-type |
- |
ISH |
 |
|
|
Lamb1 |
 |
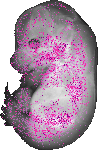 |
 |
TS23 |
14.5 dpc |
0.147 |
| Go to annotation details |
 |
 |
midbrain meninges |
 |
telencephalon meninges |
 |
lung |
|
 |
metanephros |
 |
lower jaw molar |
 |
vibrissa |
 |
cochlea |
 |
midgut |
 |
sublingual gland primordium |
 |
upper jaw molar |
 |
diencephalon meninges |
 |
spinal cord meninges |
 |
pharyngo-tympanic tube |
 |
submandibular gland primordium |
 |
urinary system |
 |
upper jaw incisor |
 |
lower jaw incisor |
 |
hindbrain meninges |
|
|
EMAGE:11988
view entry |
wild-type |
- |
ISH |
 |
|
|
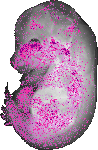
Parva |
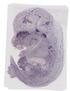 |
 |
 |
TS23 |
14.5 dpc |
0.147 |
- |
EMAGE:9664
view entry |
wild-type |
- |
ISH |
 |
|
|
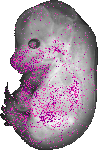
Cyld |
 |
 |
 |
TS23 |
14.5 dpc |
0.144 |
| Go to annotation details |
 |
 |
vibrissa |
 |
pituitary gland |
 |
esophagus |
|
 |
stomach |
 |
midgut |
 |
foregut-midgut junction |
 |
bladder |
 |
lung |
 |
oral epithelium |
 |
nasal cavity olfactory epithelium |
 |
naris |
 |
epidermis |
 |
rectum |
 |
hindgut |
 |
urethra of male |
|
|
EMAGE:14014
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
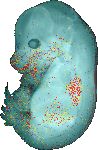 download query region as wlz file
download query region as wlz file