|
|
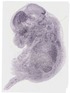
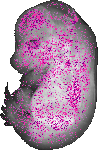
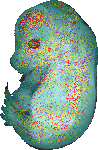
Nt5dc2 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
medulla oblongata basal plate mantle layer |
 |
thyroid gland |
 |
maxilla |
|
 |
orbito-sphenoid |
 |
lateral ventricle choroid plexus |
 |
midbrain ventricular layer |
 |
thymus primordium |
 |
bladder |
 |
pancreas |
 |
mandible |
 |
submandibular gland primordium |
 |
axial musculature |
 |
pons mantle layer |
 |
neural retina |
 |
telencephalon ventricular layer |
 |
renal cortex |
 |
lung |
 |
vault of skull |
 |
viscerocranium |
|
|
EMAGE:8684
view entry |
wild-type |
- |
ISH |
 |
|
|
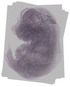
6430706D22Rik |
 |
 |
 |
TS23 |
14.5 dpc |
0.391 |
- |
EMAGE:19467
view entry |
wild-type |
- |
ISH |
 |
|
|
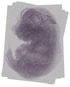
Hjurp |
 |
 |
 |
TS23 |
14.5 dpc |
0.391 |
- |
EMAGE:19468
view entry |
wild-type |
- |
ISH |
 |
|
|
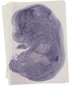
Bud31 |
 |
 |
 |
TS23 |
14.5 dpc |
0.388 |
- |
EMAGE:11456
view entry |
wild-type |
- |
ISH |
 |
|
|
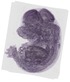
Cacybp |
 |
 |
 |
TS23 |
14.5 dpc |
0.388 |
| Go to annotation details |
 |
 |
maxilla |
 |
facial VII ganglion |
 |
midbrain |
|
 |
lower jaw molar |
 |
orbito-sphenoid |
 |
liver right lobe |
 |
thoracic ganglion |
 |
trigeminal V ganglion |
 |
hindbrain |
 |
dorsal root ganglion |
 |
vibrissa |
 |
spinal cord |
 |
nasal cavity olfactory epithelium |
 |
brain |
 |
renal cortex |
 |
glossopharyngeal IX ganglion |
 |
forebrain |
 |
submandibular gland primordium |
 |
liver left lobe |
 |
upper jaw molar |
 |
telencephalon |
 |
cervical ganglion |
 |
diencephalon |
 |
cervico-thoracic ganglion |
 |
vertebral axis musculature |
 |
mandible |
 |
thymus primordium |
|
|
EMAGE:19732
view entry |
wild-type |
- |
ISH |
 |
|
|
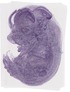
Snrpb2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.388 |
|
EMAGE:10965
view entry |
wild-type |
- |
ISH |
 |
|
|
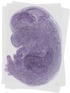
Lims1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.387 |
| Go to annotation details |
 |
 |
bladder |
 |
lower jaw molar |
 |
diencephalon lateral wall ventricular layer |
|
 |
lower jaw incisor |
 |
liver and biliary system |
 |
upper jaw molar |
 |
gut |
 |
vibrissa |
 |
midbrain ventricular layer |
 |
lung |
 |
metanephros |
 |
upper jaw incisor |
 |
liver |
 |
4th ventricle |
 |
heart |
 |
vertebral axis musculature |
 |
telencephalon ventricular layer |
 |
tongue |
 |
stomach |
 |
submandibular gland primordium |
|
|
EMAGE:11017
view entry |
wild-type |
- |
ISH |
 |
|
|
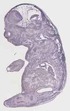
Ptrh1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.387 |
|
EMAGE:17594
view entry |
wild-type |
- |
ISH |
 |
|
|
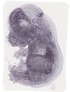
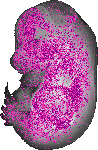
Tyms |
 |
 |
 |
TS23 |
14.5 dpc |
0.386 |
| Go to annotation details |
 |
 |
midbrain ventricular layer |
 |
diencephalon lateral wall ventricular layer |
 |
medulla oblongata alar plate ventricular layer |
|
 |
body cavity or lining |
 |
integumental system |
 |
gland |
 |
tail |
 |
mesenchyme |
 |
telencephalon ventricular layer |
 |
visceral organ system |
 |
vertebral axis musculature |
 |
sensory organ system |
 |
cardiovascular system |
 |
rest of cerebellum marginal layer |
 |
limb |
|
|
EMAGE:17349
view entry |
wild-type |
- |
ISH |
 |
|
|
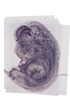
Sfrp1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.386 |
- |
EMAGE:9316
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file