|
|
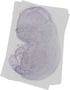
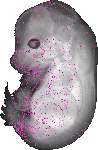
Elmo3 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
nasal cavity respiratory epithelium |
 |
stomach |
 |
nucleus pulposus |
|
 |
pharynx epithelium |
 |
rectum |
 |
midgut |
 |
humerus |
 |
oral epithelium |
 |
rib |
 |
femur |
 |
thymus primordium |
 |
liver |
 |
tongue epithelium |
 |
Meckel's cartilage |
 |
lung |
 |
nasal cavity olfactory epithelium |
|
|
EMAGE:14291
view entry |
wild-type |
- |
ISH |
 |
|
|
Olfr70 |
 |
 |
 |
TS23 |
14.5 dpc |
0.121 |
|
EMAGE:12776
view entry |
wild-type |
- |
ISH |
 |
|
|
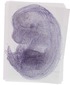
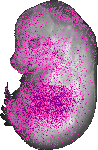
Mir187 |
 |
 |
 |
TS23 |
14.5 dpc |
0.118 |
|
EMAGE:13745
view entry |
wild-type |
- |
ISH |
 |
|
|
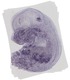
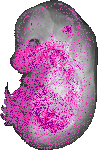
Pla2g4a |
 |
 |
 |
TS23 |
14.5 dpc |
0.117 |
| Go to annotation details |
 |
 |
viscerocranium |
 |
epidermis |
 |
nasal septum |
|
 |
bladder |
 |
kidney calyx |
 |
kidney pelvis |
 |
hindgut |
 |
rectum |
 |
midgut |
 |
vibrissa |
 |
4th ventricle lateral recess |
 |
thymus primordium |
 |
head mesenchyme |
 |
submandibular gland primordium |
 |
urethra of male |
 |
thyroid cartilage |
 |
trachea |
 |
pharyngo-tympanic tube |
 |
lateral ventricle choroid plexus |
|
|
EMAGE:11157
view entry |
wild-type |
- |
ISH |
 |
|
|
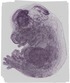
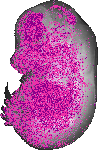
Cdca3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.116 |
| Go to annotation details |
 |
 |
liver right lobe |
 |
midbrain ventricular layer |
 |
diencephalon lateral wall ventricular layer |
|
 |
rest of cerebellum marginal layer |
 |
submandibular gland primordium |
 |
liver left lobe |
 |
mesenchyme |
 |
nasal cavity olfactory epithelium |
 |
telencephalon ventricular layer |
 |
olfactory cortex ventricular layer |
 |
renal cortex |
 |
orbito-sphenoid |
 |
rest of cerebellum ventricular layer |
 |
thymus primordium |
 |
ventral grey horn |
 |
hypothalamus ventricular layer |
|
|
EMAGE:19477
view entry |
wild-type |
- |
ISH |
 |
|
|
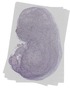
Zfp747 |
 |
 |
 |
TS23 |
14.5 dpc |
0.116 |
- |
EMAGE:14290
view entry |
wild-type |
- |
ISH |
 |
|
|
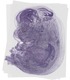
Bub1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.114 |
| Go to annotation details |
 |
 |
naris |
 |
metanephros |
 |
genital tubercle of male |
|
 |
adrenal gland |
 |
lower jaw incisor |
 |
upper jaw molar |
 |
cochlear duct |
 |
testis |
 |
submandibular gland primordium |
 |
lower jaw molar |
 |
midgut |
 |
4th ventricle |
 |
vibrissa |
 |
tongue |
 |
thymus primordium |
 |
lung |
 |
upper jaw incisor |
 |
renal cortex |
 |
liver |
 |
midgut loop |
 |
pancreas |
 |
viscerocranium |
 |
telencephalon ventricular layer |
 |
rib |
 |
cochlea |
 |
3rd ventricle |
 |
retina |
 |
foregut-midgut junction |
|
|
EMAGE:10865
view entry |
wild-type |
- |
ISH |
 |
|
|
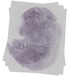
Tfam |
 |
 |
 |
TS23 |
14.5 dpc |
0.113 |
|
EMAGE:19369
view entry |
wild-type |
- |
ISH |
 |
|
|
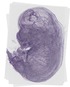
Nfatc4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.113 |
- |
EMAGE:21026
view entry |
wild-type |
- |
ISH |
 |
|
|
Prss29 |
 |
 |
 |
TS23 |
14.5 dpc |
0.113 |
- |
EMAGE:18604
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file