|
|
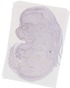
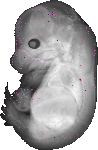
Gm7254 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:13273
view entry |
wild-type |
- |
ISH |
 |
|
|
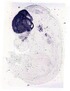
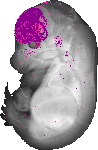
Foxg1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.027 |
- |
EMAGE:9057
view entry |
wild-type |
- |
ISH |
 |
|
|
Tha1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.027 |
- |
EMAGE:21027
view entry |
wild-type |
- |
ISH |
 |
|
|
Marveld2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.027 |
| Go to annotation details |
 |
 |
pancreas |
 |
kidney calyx |
 |
midgut |
|
 |
nasal capsule |
 |
foregut-midgut junction |
 |
stomach |
 |
hindgut |
 |
kidney pelvis |
 |
bladder |
|
|
EMAGE:11045
view entry |
wild-type |
- |
ISH |
 |
|
|
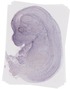
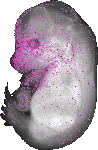
Amdhd2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.025 |
|
EMAGE:11242
view entry |
wild-type |
- |
ISH |
 |
|
|
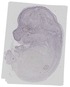
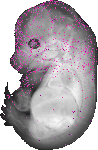
Atp11c |
 |
 |
 |
TS23 |
14.5 dpc |
0.024 |
| Go to annotation details |
 |
choroid invagination |
 |
diencephalon roof plate |
 |
metencephalon part of 4th ventricle choroid plexus |
|
|
|
|
EMAGE:15119
view entry |
wild-type |
- |
ISH |
 |
|
|
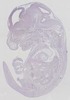
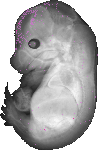
Fosl1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.024 |
- |
EMAGE:11092
view entry |
wild-type |
- |
ISH |
 |
|
|
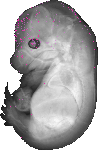
Samsn1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.024 |
- |
EMAGE:15343
view entry |
wild-type |
- |
ISH |
 |
|
|
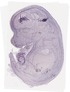
Folr1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.024 |
| Go to annotation details |
 |
 |
midgut |
 |
4th ventricle |
 |
foregut-midgut junction |
|
 |
lateral ventricle choroid plexus |
 |
hindgut |
 |
kidney calyx |
|
|
EMAGE:10556
view entry |
wild-type |
- |
ISH |
 |
|
|
Gcg |
 |
 |
 |
TS23 |
14.5 dpc |
0.023 |
|
EMAGE:12640
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file