|
|
Rnaset2b |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
lateral ventricle choroid plexus |
 |
metencephalon floor plate |
 |
diencephalon roof plate |
|
 |
thyroid gland |
 |
medulla oblongata floor plate |
 |
ovary |
 |
pancreas |
 |
vibrissa |
 |
adrenal gland |
 |
mandible |
 |
thymus primordium |
 |
metanephros |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
midbrain roof plate |
 |
orbito-sphenoid |
|
|
EMAGE:20505
view entry |
wild-type |
- |
ISH |
 |
|
|
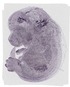
Rnaset2a |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
orbito-sphenoid |
 |
diencephalon roof plate |
|
 |
vibrissa |
 |
thymus primordium |
 |
lateral ventricle choroid plexus |
 |
adrenal gland |
 |
medulla oblongata floor plate |
 |
metencephalon floor plate |
 |
midbrain roof plate |
 |
mandible |
 |
pancreas |
 |
ovary |
 |
thyroid gland |
 |
metanephros |
|
|
EMAGE:20506
view entry |
wild-type |
- |
ISH |
 |
|
|
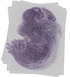
Emg1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.557 |
- |
EMAGE:19368
view entry |
wild-type |
- |
ISH |
 |
|
|
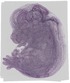
Slc39a1 |
 |
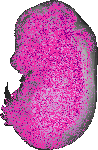 |
 |
TS23 |
14.5 dpc |
0.551 |
| Go to annotation details |
 |
 |
olfactory cortex ventricular layer |
 |
spinal cord ventricular layer |
 |
hypothalamus ventricular layer |
|
 |
thymus primordium |
 |
rest of cerebellum ventricular layer |
 |
diencephalon lateral wall ventricular layer |
 |
medulla oblongata basal plate ventricular layer |
 |
pons ventricular layer |
 |
telencephalon ventricular layer |
 |
midbrain ventricular layer |
|
|
EMAGE:19475
view entry |
wild-type |
- |
ISH |
 |
|
|
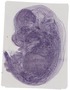
Grrp1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.550 |
| Go to annotation details |
 |
 |
vault of skull |
 |
upper jaw molar |
 |
diencephalon lateral wall ventricular layer |
|
 |
shoulder rest of mesenchyme |
 |
hypothalamus ventricular layer |
 |
clavicle |
 |
lower leg rest of mesenchyme |
 |
rest of cerebellum ventricular layer |
 |
cornea epithelium |
 |
ventral grey horn |
 |
pancreas |
 |
nasal cavity respiratory epithelium |
 |
vertebral axis musculature |
 |
thyroid gland |
 |
telencephalon ventricular layer |
 |
upper jaw incisor |
 |
mandible |
 |
pituitary gland |
 |
olfactory cortex ventricular layer |
 |
medulla oblongata basal plate ventricular layer |
 |
tongue muscle |
 |
midbrain ventricular layer |
 |
naris |
 |
tail mesenchyme |
 |
maxilla |
 |
upper leg muscle |
 |
thymus primordium |
 |
orbito-sphenoid |
 |
testis |
 |
pons ventricular layer |
 |
nasal cavity olfactory epithelium |
 |
lower jaw incisor |
 |
lower jaw molar |
 |
upper arm muscle |
|
|
EMAGE:9934
view entry |
wild-type |
- |
ISH |
 |
|
|
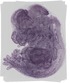
Nop2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.550 |
| Go to annotation details |
 |
 |
upper jaw incisor |
 |
renal cortex |
 |
liver lobe |
|
 |
submandibular gland primordium |
 |
lower jaw molar |
 |
vibrissa |
 |
upper jaw molar |
 |
thymus primordium |
 |
vertebral axis musculature |
 |
nasal cavity olfactory epithelium |
 |
olfactory cortex ventricular layer |
 |
lower jaw incisor |
 |
midbrain ventricular layer |
 |
telencephalon ventricular layer |
|
|
EMAGE:19479
view entry |
wild-type |
- |
ISH |
 |
|
|
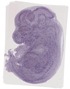
Sec11a |
 |
 |
 |
TS23 |
14.5 dpc |
0.549 |
| Go to annotation details |
 |
 |
trigeminal V ganglion |
 |
clavicle |
 |
vagus X ganglion |
|
 |
thoracic ganglion |
 |
pancreas |
 |
basisphenoid bone |
 |
ventral grey horn |
 |
submandibular gland primordium |
 |
vibrissa |
 |
upper jaw incisor |
 |
nasal cavity respiratory epithelium |
 |
pons mantle layer |
 |
dorsal root ganglion |
 |
kidney calyx |
 |
medulla oblongata basal plate mantle layer |
 |
lower jaw incisor |
 |
cervico-thoracic ganglion |
 |
glossopharyngeal IX ganglion |
 |
testis |
 |
cervical ganglion |
 |
orbito-sphenoid |
 |
nasal cavity olfactory epithelium |
 |
Meckel's cartilage |
|
|
EMAGE:10849
view entry |
wild-type |
- |
ISH |
 |
|
|
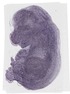
Timm9 |
 |
 |
 |
TS23 |
14.5 dpc |
0.548 |
| Go to annotation details |
 |
 |
otic capsule |
 |
renal cortex |
 |
telencephalon ventricular layer |
|
 |
olfactory cortex ventricular layer |
 |
orbito-sphenoid |
|
|
EMAGE:17157
view entry |
wild-type |
- |
ISH |
 |
|
|
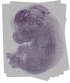
Hirip3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.548 |
- |
EMAGE:19704
view entry |
wild-type |
- |
ISH |
 |
|
|
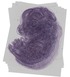
Sec61g |
 |
 |
 |
TS23 |
14.5 dpc |
0.548 |
- |
EMAGE:19365
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
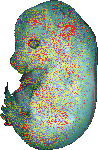 download query region as wlz file
download query region as wlz file