|
|
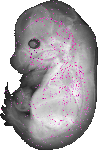
Rapgef4 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:16308
view entry |
wild-type |
- |
ISH |
 |
|
|
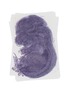
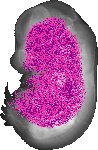
Nup93 |
 |
 |
 |
TS23 |
14.5 dpc |
0.109 |
- |
EMAGE:17340
view entry |
wild-type |
- |
ISH |
 |
|
|
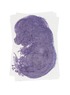
Zmynd19 |
 |
 |
 |
TS23 |
14.5 dpc |
0.108 |
- |
EMAGE:17341
view entry |
wild-type |
- |
ISH |
 |
|
|
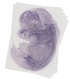
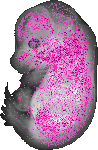
Prokr1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.100 |
| Go to annotation details |
 |
 |
neural retina |
 |
dorsal root ganglion |
 |
medulla oblongata alar plate ventricular layer |
|
 |
telencephalon ventricular layer |
 |
midbrain ventricular layer |
 |
trigeminal V ganglion |
 |
glossopharyngeal IX ganglion |
 |
spinal cord ventricular layer |
 |
mesenchyme |
 |
diencephalon lateral wall ventricular layer |
 |
medulla oblongata basal plate ventricular layer |
|
|
EMAGE:12236
view entry |
wild-type |
- |
ISH |
 |
|
|
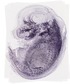
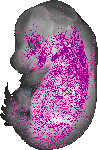
Car3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.100 |
| Go to annotation details |
 |
 |
shoulder mesenchyme |
 |
axial skeleton tail region |
 |
vertebral axis musculature |
|
 |
mandible |
 |
lower leg mesenchyme |
 |
nucleus pulposus |
 |
maxilla |
 |
esophagus |
 |
upper arm mesenchyme |
 |
forearm mesenchyme |
 |
bladder |
 |
upper leg mesenchyme |
 |
tail mesenchyme |
 |
cranium |
 |
rib |
 |
foot mesenchyme |
 |
male reproductive system |
 |
kidney pelvis |
|
|
EMAGE:7147
view entry |
wild-type |
- |
ISH |
 |
|
|
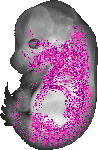
Itgb1bp2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.099 |
| Go to annotation details |
 |
 |
tongue muscle |
 |
knee rest of mesenchyme |
 |
lower leg rest of mesenchyme |
|
 |
elbow rest of mesenchyme |
 |
vertebral axis musculature |
 |
tail paraxial mesenchyme |
 |
forearm rest of mesenchyme |
 |
hip rest of mesenchyme |
 |
hand mesenchyme |
 |
upper arm muscle |
 |
upper leg muscle |
 |
upper arm rest of mesenchyme |
 |
tarsus rest of mesenchyme |
 |
shoulder rest of mesenchyme |
 |
diaphragm |
|
|
EMAGE:11539
view entry |
wild-type |
- |
ISH |
 |
|
|
Phlda3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.099 |
- |
EMAGE:20019
view entry |
wild-type |
- |
ISH |
 |
|
|
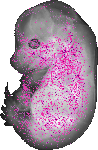
Elf4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.099 |
| Go to annotation details |
 |
 |
hypothalamus mantle layer |
 |
pancreas |
 |
mesenchyme |
|
 |
bladder |
 |
vibrissa |
 |
midgut |
 |
telencephalon mantle layer |
 |
maxilla |
 |
pericardium |
 |
mandible |
 |
lung |
 |
hindgut |
 |
stomach |
|
|
EMAGE:21713
view entry |
wild-type |
- |
ISH |
 |
|
|
Scara3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.099 |
|
EMAGE:14343
view entry |
wild-type |
- |
ISH |
 |
|
|
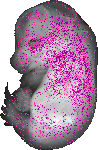
Nid1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.098 |
| Go to annotation details |
 |
 |
viscerocranium |
 |
vertebral axis musculature |
 |
extrinsic ocular muscle |
|
 |
trigeminal V nerve maxillary division |
 |
diencephalon meninges |
 |
medulla oblongata basal plate ventricular layer |
 |
telencephalon meninges |
 |
pons ventricular layer |
 |
rest of cerebellum ventricular layer |
 |
spinal cord meninges |
 |
diaphragm |
 |
axial skeleton cervical region |
 |
lens |
 |
vibrissa |
 |
hindbrain meninges |
 |
dorsal grey horn |
 |
midbrain meninges |
|
|
EMAGE:8607
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file