|
|
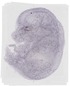
Hsd17b11 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
|
EMAGE:20334
view entry |
wild-type |
- |
ISH |
 |
|
|
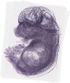
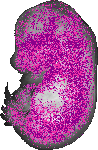
Islr |
 |
 |
 |
TS23 |
14.5 dpc |
0.192 |
| Go to annotation details |
 |
 |
hindbrain meninges |
 |
respiratory system |
 |
reproductive system |
|
 |
diencephalon meninges |
 |
urinary system |
 |
limb |
 |
integumental system |
 |
mesenchyme |
 |
spinal cord meninges |
 |
body cavity or lining |
 |
alimentary system |
 |
vertebral axis musculature |
 |
gland |
 |
telencephalon meninges |
 |
midbrain meninges |
|
|
EMAGE:20020
view entry |
wild-type |
- |
ISH |
 |
|
|
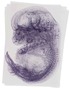
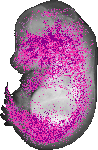
Kctd12 |
 |
 |
 |
TS23 |
14.5 dpc |
0.191 |
- |
EMAGE:12003
view entry |
wild-type |
- |
ISH |
 |
|
|
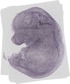
Lamp1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.191 |
| Go to annotation details |
 |
 |
diencephalon lateral wall ventricular layer |
 |
dorsal root ganglion |
 |
midbrain ventricular layer |
|
 |
midbrain floor plate |
 |
metencephalon floor plate |
 |
mandible |
 |
medulla oblongata basal plate ventricular layer |
 |
trigeminal V ganglion |
 |
medulla oblongata floor plate |
 |
orbito-sphenoid |
 |
neural retina |
 |
maxilla |
 |
sternum |
 |
pons ventricular layer |
 |
ventral grey horn |
 |
glossopharyngeal IX ganglion |
|
|
EMAGE:19072
view entry |
wild-type |
- |
ISH |
 |
|
|
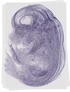
Emilin1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.190 |
|
EMAGE:10927
view entry |
wild-type |
- |
ISH |
 |
|
|
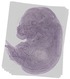
Trabd |
 |
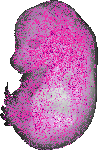 |
 |
TS23 |
14.5 dpc |
0.190 |
- |
EMAGE:19499
view entry |
wild-type |
- |
ISH |
 |
|
|
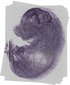
Nfix |
 |
 |
 |
TS23 |
14.5 dpc |
0.190 |
- |
EMAGE:19386
view entry |
wild-type |
- |
ISH |
 |
|
|
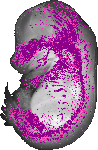
Lect1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.190 |
| Go to annotation details |
 |
 |
cochlea |
 |
hindlimb digit 2 mesenchyme |
 |
rib |
|
 |
nasal septum mesenchyme |
 |
leg |
 |
hindlimb digit 4 mesenchyme |
 |
pectoral girdle and thoracic body wall skeleton |
 |
axial skeleton |
 |
hindlimb digit 5 mesenchyme |
 |
hand mesenchyme |
 |
axial skeleton tail region |
 |
viscerocranium |
 |
orbito-sphenoid |
 |
hindlimb digit 1 mesenchyme |
 |
vertebral axis musculature |
 |
chondrocranium |
 |
cochlear duct mesenchyme |
 |
hindlimb digit 3 mesenchyme |
|
|
EMAGE:19343
view entry |
wild-type |
- |
ISH |
 |
|
|
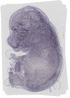
Metrn |
 |
 |
 |
TS23 |
14.5 dpc |
0.190 |
| Go to annotation details |
 |
 |
telencephalon ventricular layer |
 |
midgut |
 |
rest of cerebellum ventricular layer |
|
 |
olfactory cortex marginal layer |
 |
hypothalamus ventricular layer |
 |
neural retina |
 |
ventral grey horn |
 |
pons ventricular layer |
 |
diencephalon lateral wall ventricular layer |
 |
olfactory cortex ventricular layer |
 |
spinal cord ventricular layer |
|
|
EMAGE:18290
view entry |
wild-type |
- |
ISH |
 |
|
|
Mir345 |
 |
 |
 |
TS23 |
14.5 dpc |
0.190 |
- |
EMAGE:19158
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file