|
|
Mrgpra2b |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:14478
view entry |
wild-type |
- |
ISH |
 |
|
|
Mrgpra2a |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:14479
view entry |
wild-type |
- |
ISH |
 |
|
|
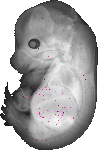
Pah |
 |
 |
 |
TS23 |
14.5 dpc |
0.014 |
|
EMAGE:10459
view entry |
wild-type |
- |
ISH |
 |
|
|
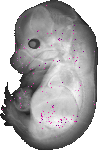
Igsf5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
|
EMAGE:10553
view entry |
wild-type |
- |
ISH |
 |
|
|
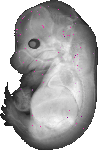
Lhcgr |
 |
 |
 |
TS23 |
14.5 dpc |
0.010 |
|
EMAGE:21065
view entry |
wild-type |
- |
ISH |
 |
|
|
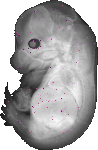
Saa4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.008 |
- |
EMAGE:13513
view entry |
wild-type |
- |
ISH |
 |
|
|
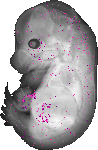
2210415F13Rik |
 |
 |
 |
TS23 |
14.5 dpc |
0.008 |
|
EMAGE:20515
view entry |
wild-type |
- |
ISH |
 |
|
|
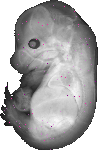
Extl1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
| Go to annotation details |
 |
 |
hindlimb digit 1 metatarsal |
 |
hindlimb digit 3 metatarsal |
 |
fibula |
|
 |
hindlimb digit 5 metatarsal |
 |
tibia |
 |
exoccipital bone |
 |
hindlimb digit 4 metatarsal |
 |
orbito-sphenoid |
 |
viscerocranium |
 |
hindlimb digit 2 metatarsal |
 |
nasal septum |
|
|
EMAGE:15016
view entry |
wild-type |
- |
ISH |
 |
|
|
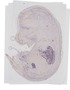
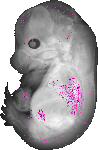
Wnt2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
|
EMAGE:21702
view entry |
wild-type |
- |
ISH |
 |
|
|
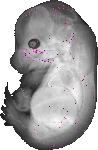
Ehf |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
| Go to annotation details |
 |
 |
sublingual gland primordium |
 |
nasal cavity olfactory epithelium |
 |
anterior naris epithelium |
|
 |
pancreas |
 |
submandibular gland primordium |
 |
metanephros |
 |
bladder |
|
|
EMAGE:18132
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
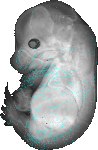 download query region as wlz file
download query region as wlz file