|
|
Taldo1 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:7298
view entry |
wild-type |
- |
ISH |
 |
|
|
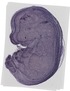
Mrps6 |
 |
 |
 |
TS23 |
14.5 dpc |
0.756 |
| Go to annotation details |
 |
 |
telencephalon |
 |
medulla oblongata part of 4th ventricle choroid plexus |
 |
lateral ventricle choroid plexus |
|
 |
metencephalon |
|
|
EMAGE:10053
view entry |
wild-type |
- |
ISH |
 |
|
|
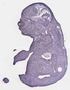
Sae1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.756 |
|
EMAGE:17634
view entry |
wild-type |
- |
ISH |
 |
|
|
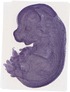
Egln2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.756 |
- |
EMAGE:12170
view entry |
wild-type |
- |
ISH |
 |
|
|
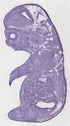
Rpl4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
- |
EMAGE:17301
view entry |
wild-type |
- |
ISH |
 |
|
|
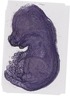
Krt4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
| Go to annotation details |
 |
 |
lower jaw molar |
 |
trachea |
 |
upper jaw molar |
|
 |
thymus primordium |
 |
oral epithelium |
 |
vibrissa |
 |
esophagus |
 |
naris |
 |
lower jaw incisor |
 |
cornea epithelium |
 |
epidermis |
 |
bladder |
 |
rectum |
 |
nasal cavity respiratory epithelium |
 |
upper jaw incisor |
 |
urethra of male |
 |
nucleus pulposus |
|
|
EMAGE:16846
view entry |
wild-type |
- |
ISH |
 |
|
|
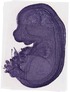
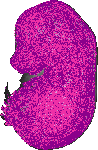
Hist3h2ba |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
|
EMAGE:12169
view entry |
wild-type |
- |
ISH |
 |
|
|
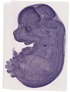
Morn4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
| Go to annotation details |
 |
 |
neural retina |
 |
tongue |
 |
trigeminal V ganglion |
|
 |
spinal cord |
 |
midgut |
 |
vestibulocochlear VIII ganglion |
 |
dorsal root ganglion |
 |
glossopharyngeal IX ganglion |
 |
cervical ganglion |
 |
cervico-thoracic ganglion |
 |
vagus X ganglion |
 |
brain |
 |
thoracic ganglion |
 |
stomach |
 |
facial VII ganglion |
 |
trigeminal V nerve |
 |
nasal cavity olfactory epithelium |
|
|
EMAGE:12171
view entry |
wild-type |
- |
ISH |
 |
|
|
Synj2bp |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
- |
EMAGE:17055
view entry |
wild-type |
- |
ISH |
 |
|
|
Tcf3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.755 |
- |
EMAGE:17265
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file