|
|
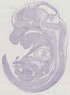
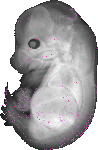
Gemin7 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:11549
view entry |
wild-type |
- |
ISH |
 |
|
|
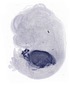
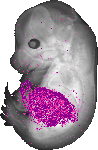
Serpina1c |
 |
 |
 |
TS23 |
14.5 dpc |
0.036 |
| Go to annotation details |
 |
 |
midgut loop lumen |
 |
liver lobe |
 |
pancreas |
|
 |
rectum lumen |
 |
kidney calyx |
 |
foregut-midgut junction lumen |
 |
midgut |
 |
rest of hindgut lumen |
|
|
EMAGE:10804
view entry |
wild-type |
- |
ISH |
 |
|
|
Pcsk9 |
 |
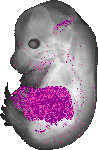 |
 |
TS23 |
14.5 dpc |
0.035 |
| Go to annotation details |
 |
 |
upper jaw incisor |
 |
rest of cerebellum marginal layer |
 |
anal canal |
|
 |
kidney calyx |
 |
hindgut |
 |
foregut-midgut junction |
 |
stomach |
 |
liver lobe |
 |
midgut |
 |
lower jaw incisor |
 |
pancreas |
|
|
EMAGE:11112
view entry |
wild-type |
- |
ISH |
 |
|
|
Gm4598 |
 |
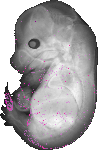 |
 |
TS23 |
14.5 dpc |
0.034 |
- |
EMAGE:15918
view entry |
wild-type |
- |
ISH |
 |
|
|
Serpina1a |
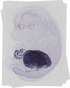 |
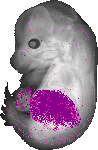 |
 |
TS23 |
14.5 dpc |
0.034 |
|
EMAGE:14172
view entry |
wild-type |
- |
ISH |
 |
|
|
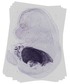
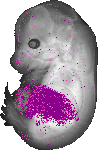
Serpina1b |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
| Go to annotation details |
 |
 |
rectum lumen |
 |
midgut loop lumen |
 |
pancreas |
|
 |
liver left lobe |
 |
liver right lobe |
 |
nucleus pulposus |
 |
kidney calyx |
|
|
EMAGE:14268
view entry |
wild-type |
- |
ISH |
 |
|
|
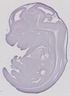
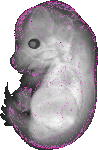
Hoxc10 |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
- |
EMAGE:10226
view entry |
wild-type |
- |
ISH |
 |
|
|
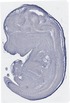
Mccc1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.034 |
- |
EMAGE:9748
view entry |
wild-type |
- |
ISH |
 |
|
|
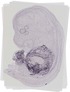
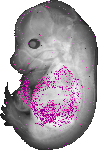
Gm98 |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
- |
EMAGE:13064
view entry |
wild-type |
- |
ISH |
 |
|
|
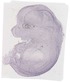
Btn1a1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
|
EMAGE:7317
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file