|
|
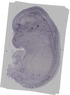
Car11 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
facial VII ganglion |
 |
rest of cerebellum mantle layer |
 |
trigeminal V ganglion |
|
 |
olfactory cortex marginal layer |
 |
telencephalon mantle layer |
 |
diencephalon lateral wall mantle layer |
 |
midbrain marginal layer |
 |
midbrain mantle layer |
 |
dorsal root ganglion |
|
|
EMAGE:15436
view entry |
wild-type |
- |
ISH |
 |
|
|
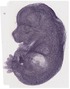
Tsnax |
 |
 |
 |
TS23 |
14.5 dpc |
0.573 |
| Go to annotation details |
 |
 |
integumental system |
 |
tail skeleton |
 |
vertebral axis musculature |
|
 |
body cavity or lining |
 |
gland |
 |
alimentary system |
 |
limb |
 |
reproductive system |
 |
nervous system |
 |
tail nervous system |
 |
cardiovascular system |
 |
urinary system |
 |
respiratory system |
 |
sensory organ system |
 |
skeleton |
 |
visceral organ system |
 |
tail |
|
|
EMAGE:7336
view entry |
wild-type |
- |
ISH |
 |
|
|
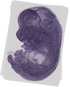
Efs |
 |
 |
 |
TS23 |
14.5 dpc |
0.572 |
- |
EMAGE:11718
view entry |
wild-type |
- |
ISH |
 |
|
|
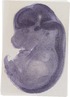
Mir466g |
 |
 |
 |
TS23 |
14.5 dpc |
0.572 |
| Go to annotation details |
 |
 |
diencephalon lateral wall mantle layer |
 |
rest of cerebellum mantle layer |
 |
spinal cord mantle layer |
|
 |
medulla oblongata basal plate mantle layer |
 |
telencephalon mantle layer |
 |
neural retina |
 |
midbrain mantle layer |
 |
dorsal root ganglion |
 |
vestibulocochlear VIII ganglion |
 |
medulla oblongata alar plate mantle layer |
 |
trigeminal V ganglion |
 |
pons marginal layer |
 |
facial VII ganglion |
 |
medulla oblongata basal plate marginal layer |
 |
vagus X ganglion |
 |
glossopharyngeal IX ganglion |
 |
pons mantle layer |
|
|
EMAGE:13871
view entry |
wild-type |
- |
ISH |
 |
|
|
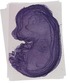
Adprhl1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.572 |
- |
EMAGE:11302
view entry |
wild-type |
- |
ISH |
 |
|
|
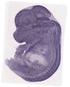
Ablim3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.571 |
| Go to annotation details |
 |
 |
telencephalon |
 |
vagus X ganglion |
 |
dorsal root ganglion |
|
 |
vestibulocochlear VIII ganglion |
 |
midbrain |
 |
hindbrain |
 |
spinal cord |
 |
facial VII ganglion |
 |
diencephalon |
 |
neural retina |
 |
glossopharyngeal IX ganglion |
 |
heart valve |
 |
trigeminal V ganglion |
 |
trigeminal V nerve |
|
|
EMAGE:12076
view entry |
wild-type |
- |
ISH |
 |
|
|
Rcbtb2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.571 |
- |
EMAGE:17695
view entry |
wild-type |
- |
ISH |
 |
|
|
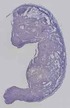
Mir706 |
 |
 |
 |
TS23 |
14.5 dpc |
0.571 |
- |
EMAGE:16282
view entry |
wild-type |
- |
ISH |
 |
|
|
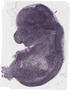
Wdr6 |
 |
 |
 |
TS23 |
14.5 dpc |
0.570 |
- |
EMAGE:17657
view entry |
wild-type |
- |
ISH |
 |
|
|
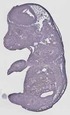
Ap2s1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.570 |
- |
EMAGE:17556
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file