|
|
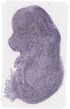
C1qtnf7 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
basioccipital bone |
 |
humerus |
 |
radius |
|
 |
tibia |
 |
temporal bone petrous part |
 |
viscerocranium |
 |
fibula |
 |
scapula |
 |
pelvic girdle skeleton |
 |
orbito-sphenoid |
 |
basisphenoid bone |
 |
ulna |
 |
exoccipital bone |
|
|
EMAGE:16787
view entry |
wild-type |
- |
ISH |
 |
|
|
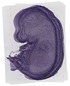
Gbp8 |
 |
 |
 |
TS23 |
14.5 dpc |
0.763 |
|
EMAGE:12381
view entry |
wild-type |
- |
ISH |
 |
|
|
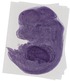
Ubac1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.763 |
| Go to annotation details |
 |
 |
vibrissa |
 |
liver left lobe |
 |
facial VII ganglion |
|
 |
trigeminal V ganglion |
 |
glossopharyngeal IX ganglion |
 |
dorsal root ganglion |
 |
liver right lobe |
 |
vestibulocochlear VIII ganglion |
|
|
EMAGE:11102
view entry |
wild-type |
- |
ISH |
 |
|
|
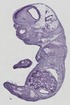
Ctsb |
 |
 |
 |
TS23 |
14.5 dpc |
0.763 |
- |
EMAGE:11101
view entry |
wild-type |
- |
ISH |
 |
|
|
Nudt3 |
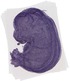 |
 |
 |
TS23 |
14.5 dpc |
0.762 |
- |
EMAGE:12344
view entry |
wild-type |
- |
ISH |
 |
|
|
Becn1 |
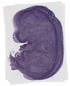 |
 |
 |
TS23 |
14.5 dpc |
0.762 |
|
EMAGE:12385
view entry |
wild-type |
- |
ISH |
 |
|
|
Orai3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.761 |
|
EMAGE:12348
view entry |
wild-type |
- |
ISH |
 |
|
|
Cct7 |
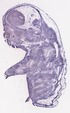 |
 |
 |
TS23 |
14.5 dpc |
0.761 |
- |
EMAGE:16546
view entry |
wild-type |
- |
ISH |
 |
|
|
Calm3 |
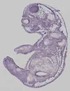 |
 |
 |
TS23 |
14.5 dpc |
0.760 |
- |
EMAGE:19449
view entry |
wild-type |
- |
ISH |
 |
|
|
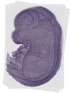
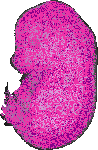
Traf7 |
 |
 |
 |
TS23 |
14.5 dpc |
0.760 |
- |
EMAGE:10870
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
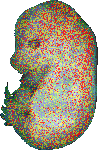 download query region as wlz file
download query region as wlz file