|
|
Lysmd2 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
|
EMAGE:20548
view entry |
wild-type |
- |
ISH |
 |
|
|
Adarb2 |
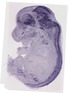 |
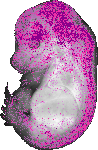 |
 |
TS23 |
14.5 dpc |
0.034 |
| Go to annotation details |
 |
 |
axial musculature |
 |
rest of cerebellum mantle layer |
 |
foot |
|
 |
spinal cord mantle layer |
 |
nasal cavity olfactory epithelium |
 |
hand |
 |
vestibulocochlear VIII ganglion |
 |
medulla oblongata alar plate mantle layer |
 |
heart ventricle |
 |
medulla oblongata basal plate mantle layer |
 |
lower leg rest of mesenchyme |
 |
glossopharyngeal IX ganglion |
 |
telencephalon mantle layer |
 |
tongue muscle |
 |
midbrain mantle layer |
 |
larynx |
 |
diencephalon lateral wall mantle layer |
 |
trigeminal V nerve |
 |
pons mantle layer |
 |
neural retina |
 |
upper arm muscle |
 |
dorsal root ganglion |
 |
upper leg muscle |
 |
trigeminal V ganglion |
 |
genital tubercle of male |
|
|
EMAGE:12629
view entry |
wild-type |
- |
ISH |
 |
|
|
Fgf15 |
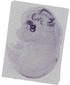 |
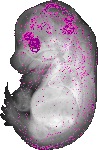 |
 |
TS23 |
14.5 dpc |
0.033 |
| Go to annotation details |
 |
 |
pons ventricular layer |
 |
olfactory cortex ventricular layer |
 |
retina nuclear layer |
|
 |
medulla oblongata basal plate ventricular layer |
 |
telencephalon mantle layer |
 |
dorsal grey horn |
 |
rest of cerebellum mantle layer |
 |
nasal cavity olfactory epithelium |
 |
diencephalon lateral wall ventricular layer |
 |
midbrain ventricular layer |
 |
medulla oblongata alar plate ventricular layer |
 |
hypothalamus ventricular layer |
 |
rest of cerebellum ventricular layer |
 |
telencephalon ventricular layer |
|
|
EMAGE:15301
view entry |
wild-type |
- |
ISH |
 |
|
|
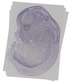
Polk |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
- |
EMAGE:15308
view entry |
wild-type |
- |
ISH |
 |
|
|
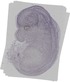
Rassf2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.033 |
| Go to annotation details |
 |
 |
midbrain meninges |
 |
pons ventricular layer |
 |
telencephalon meninges |
|
 |
spinal cord ventricular layer |
 |
midbrain ventricular layer |
 |
hindbrain meninges |
 |
medulla oblongata basal plate ventricular layer |
 |
heart valve |
 |
spinal cord meninges |
 |
diencephalon meninges |
|
|
EMAGE:15309
view entry |
wild-type |
- |
ISH |
 |
|
|
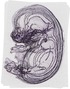
Krt5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.032 |
| Go to annotation details |
 |
 |
tongue epithelium |
 |
upper jaw molar |
 |
upper jaw incisor |
|
 |
thymus primordium |
 |
lower jaw incisor |
 |
physiological umbilical hernia epidermis |
 |
pharynx |
 |
bladder |
 |
conjunctival sac |
 |
nasal cavity respiratory epithelium |
 |
esophagus |
 |
lower jaw molar |
 |
naso-lacrimal duct |
 |
vibrissa |
 |
nasal cavity olfactory epithelium |
 |
pharyngo-tympanic tube |
 |
trachea |
 |
oral epithelium |
 |
urinary system |
|
|
EMAGE:19397
view entry |
wild-type |
- |
ISH |
 |
|
|
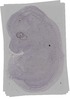
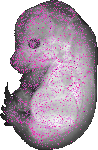
Ubn1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.032 |
- |
EMAGE:15653
view entry |
wild-type |
- |
ISH |
 |
|
|
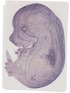
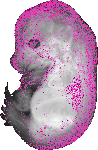
Ager |
 |
 |
 |
TS23 |
14.5 dpc |
0.032 |
|
EMAGE:11492
view entry |
wild-type |
- |
ISH |
 |
|
|
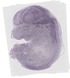
Pip5k1c |
 |
 |
 |
TS23 |
14.5 dpc |
0.032 |
| Go to annotation details |
 |
 |
facial VII ganglion |
 |
cervical ganglion |
 |
tongue |
|
 |
spinal cord |
 |
trigeminal V nerve |
 |
vagus X ganglion |
 |
brain |
 |
neural retina |
 |
thoracic ganglion |
 |
vomeronasal organ |
 |
cervico-thoracic ganglion |
 |
trigeminal V ganglion |
 |
heart ventricle |
 |
dorsal root ganglion |
 |
glossopharyngeal IX ganglion |
 |
vestibulocochlear VIII ganglion |
 |
nasal cavity olfactory epithelium |
|
|
EMAGE:20166
view entry |
wild-type |
- |
ISH |
 |
|
|
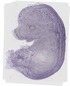
Bcl2l2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.032 |
- |
EMAGE:20147
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
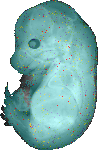 download query region as wlz file
download query region as wlz file