Reconstruction of Serial Sections
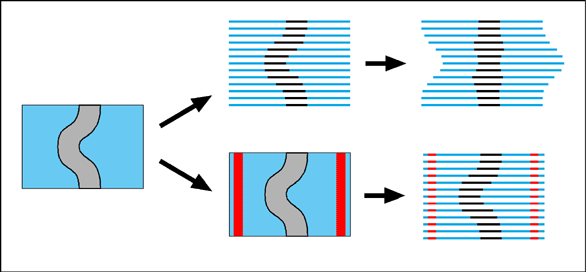
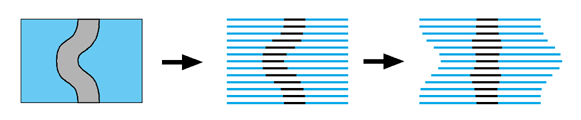
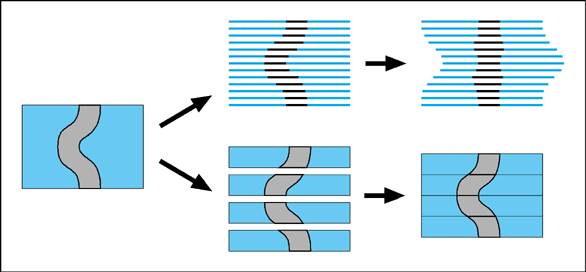
Serial sections The principle of using serial sections to build-up a reconstruction of biological tissue has been around a long time. However, it suffers from many drawbacks. Apart from the simple problem that it takes a long time to cut hundreds of sections, mount them on slides, stain them, photograph them and then process them through a computer (while trying not to loose any of them), there is also a theoretical difficulty in knowing how to fir them back together within the 3-D reconstruction. One common approach is simply too match up the structures on adjacent sections with each other. This will always suffer from the problem that structures will tend to become straightened-out along the axis perpendicular to the sections. This is illustrated by an exaggerated example in the following figure.
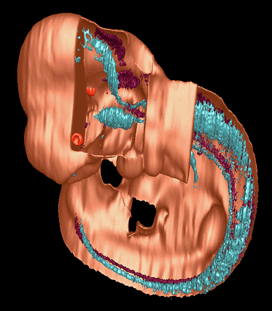
One solution to this problem is to use thick sections which retain some 3-D information about the orientation of structures which pass through the section. Thick sections like these can be scanned using a confocal microscope, and reconstructed as shown below.
An alternative solution which uses thin sections is to provide external registration markers. In one technique (Streicher et al., 2000) the specimen is embedded in a resin block which is not dissolved away after the sections have been mounted. Before sectioning, vertical holes are drilled into the block around the specimen. These holes can be seen adjacent to the tissue in the mounted sections, so that a reconstruction based on these, rather than on the tissue itself, will retain its true 3-D shape.