|
|
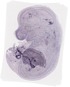
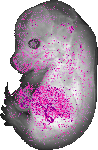
Lgals3bp |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
|
EMAGE:10595
view entry |
wild-type |
- |
ISH |
 |
|
|
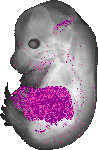
Pcsk9 |
 |
 |
 |
TS23 |
14.5 dpc |
0.353 |
| Go to annotation details |
 |
 |
liver lobe |
 |
upper jaw incisor |
 |
kidney calyx |
|
 |
midgut |
 |
anal canal |
 |
hindgut |
 |
pancreas |
 |
rest of cerebellum marginal layer |
 |
lower jaw incisor |
 |
foregut-midgut junction |
 |
stomach |
|
|
EMAGE:11112
view entry |
wild-type |
- |
ISH |
 |
|
|
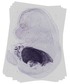
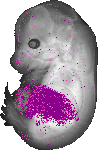
Serpina1b |
 |
 |
 |
TS23 |
14.5 dpc |
0.336 |
| Go to annotation details |
 |
 |
liver left lobe |
 |
rectum lumen |
 |
kidney calyx |
|
 |
nucleus pulposus |
 |
liver right lobe |
 |
pancreas |
 |
midgut loop lumen |
|
|
EMAGE:14268
view entry |
wild-type |
- |
ISH |
 |
|
|
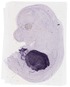
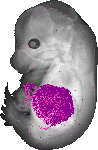
Serpina1e |
 |
 |
 |
TS23 |
14.5 dpc |
0.335 |
|
EMAGE:12543
view entry |
wild-type |
- |
ISH |
 |
|
|
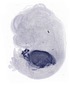
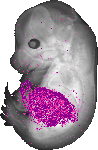
Serpina1c |
 |
 |
 |
TS23 |
14.5 dpc |
0.334 |
| Go to annotation details |
 |
 |
foregut-midgut junction lumen |
 |
pancreas |
 |
kidney calyx |
|
 |
midgut |
 |
liver lobe |
 |
rest of hindgut lumen |
 |
rectum lumen |
 |
midgut loop lumen |
|
|
EMAGE:10804
view entry |
wild-type |
- |
ISH |
 |
|
|
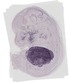
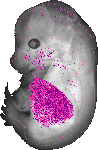
Acp5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.333 |
|
EMAGE:10955
view entry |
wild-type |
- |
ISH |
 |
|
|
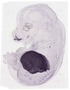
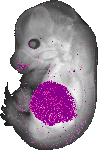
Alb |
 |
 |
 |
TS23 |
14.5 dpc |
0.333 |
| Go to annotation details |
 |
 |
liver |
 |
anterior abdominal wall musculature |
 |
lower lip |
|
 |
pectoral girdle and thoracic body wall musculature |
 |
upper lip |
|
|
EMAGE:7464
view entry |
wild-type |
- |
ISH |
 |
|
|
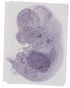
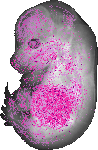
Pabpc6 |
 |
 |
 |
TS23 |
14.5 dpc |
0.332 |
| Go to annotation details |
 |
 |
upper jaw molar |
 |
vomeronasal organ |
 |
midbrain ventricular layer |
|
 |
pituitary gland |
 |
tongue muscle |
 |
maxilla |
 |
stomach |
 |
vibrissa |
 |
urethra of male |
 |
naris |
 |
lower jaw molar |
 |
hindgut |
 |
lung |
 |
esophagus |
 |
telencephalon ventricular layer |
 |
upper jaw incisor |
 |
submandibular gland primordium |
 |
thymus primordium |
 |
pancreas |
 |
mandible |
 |
lower jaw incisor |
 |
renal cortex |
 |
nasal cavity olfactory epithelium |
 |
thyroid gland |
 |
vertebral axis musculature |
 |
midgut |
 |
liver lobe |
 |
rectum |
|
|
EMAGE:11986
view entry |
wild-type |
- |
ISH |
 |
|
|
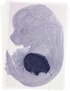
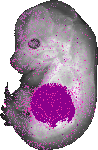
Serpina6 |
 |
 |
 |
TS23 |
14.5 dpc |
0.331 |
|
EMAGE:11556
view entry |
wild-type |
- |
ISH |
 |
|
|
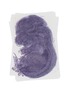
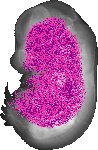
Nup93 |
 |
 |
 |
TS23 |
14.5 dpc |
0.326 |
- |
EMAGE:17340
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
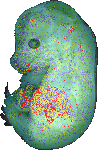 download query region as wlz file
download query region as wlz file