|
|
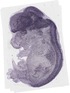
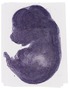
Tspan13 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
| Go to annotation details |
 |
 |
urethra of male |
 |
dorsal root ganglion |
 |
brain |
|
 |
male reproductive system |
 |
rectum |
 |
nasal cavity olfactory epithelium |
 |
bladder |
 |
cervico-thoracic ganglion |
 |
vagus X ganglion |
 |
trigeminal V ganglion |
 |
physiological umbilical hernia |
 |
cervical ganglion |
 |
vestibulocochlear VIII ganglion |
 |
stomach |
 |
thoracic ganglion |
 |
left lung |
 |
glossopharyngeal IX ganglion |
 |
facial VII ganglion |
 |
neural retina |
 |
right lung |
 |
spinal cord |
 |
thymus primordium |
|
|
EMAGE:17451
view entry |
wild-type |
- |
ISH |
 |
|
|
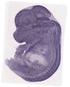
Ablim3 |
 |
 |
 |
TS23 |
14.5 dpc |
0.651 |
| Go to annotation details |
 |
 |
glossopharyngeal IX ganglion |
 |
dorsal root ganglion |
 |
telencephalon |
|
 |
neural retina |
 |
spinal cord |
 |
midbrain |
 |
trigeminal V ganglion |
 |
heart valve |
 |
vagus X ganglion |
 |
vestibulocochlear VIII ganglion |
 |
hindbrain |
 |
facial VII ganglion |
 |
diencephalon |
 |
trigeminal V nerve |
|
|
EMAGE:12076
view entry |
wild-type |
- |
ISH |
 |
|
|
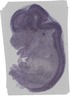
Grik5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.650 |
| Go to annotation details |
 |
 |
spinal cord |
 |
trigeminal V nerve |
 |
trigeminal V ganglion |
|
 |
facial VII ganglion |
 |
midbrain |
 |
glossopharyngeal IX ganglion |
 |
vagus X ganglion |
 |
vibrissa |
 |
forebrain |
 |
vestibulocochlear VIII ganglion |
 |
hindbrain |
 |
dorsal root ganglion |
 |
neural retina |
 |
lung |
|
|
EMAGE:15287
view entry |
wild-type |
- |
ISH |
 |
|
|
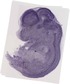
Drd5 |
 |
 |
 |
TS23 |
14.5 dpc |
0.650 |
| Go to annotation details |
 |
 |
trigeminal V ganglion |
 |
dorsal root ganglion |
 |
vestibulocochlear VIII ganglion |
|
 |
neural retina |
 |
facial VII ganglion |
 |
glossopharyngeal IX ganglion |
 |
vagus X ganglion |
|
|
EMAGE:11831
view entry |
wild-type |
- |
ISH |
 |
|
|
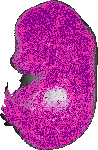
Wnt7a |
 |
 |
 |
TS23 |
14.5 dpc |
0.650 |
- |
EMAGE:19714
view entry |
wild-type |
- |
ISH |
 |
|
|
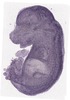
Wdr54 |
 |
 |
 |
TS23 |
14.5 dpc |
0.649 |
|
EMAGE:7791
view entry |
wild-type |
- |
ISH |
 |
|
|
Trib2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.649 |
- |
EMAGE:18372
view entry |
wild-type |
- |
ISH |
 |
|
|
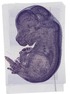
Enc1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.649 |
- |
EMAGE:9282
view entry |
wild-type |
- |
ISH |
 |
|
|
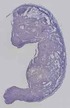
Mir706 |
 |
 |
 |
TS23 |
14.5 dpc |
0.648 |
- |
EMAGE:16282
view entry |
wild-type |
- |
ISH |
 |
|
|
Map3k12 |
 |
 |
 |
TS23 |
14.5 dpc |
0.648 |
- |
EMAGE:17233
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
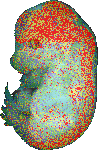 download query region as wlz file
download query region as wlz file