|
|
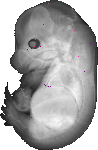
Zfp316 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:8490
view entry |
wild-type |
- |
ISH |
 |
|
|
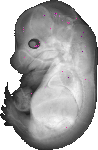
Plcl1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.016 |
| Go to annotation details |
 |
 |
vibrissa |
 |
midbrain floor plate |
 |
olfactory cortex mantle layer |
|
 |
pons mantle layer |
 |
medulla oblongata basal plate mantle layer |
 |
medulla oblongata alar plate mantle layer |
 |
telencephalon mantle layer |
 |
ventral grey horn |
 |
diencephalon lateral wall mantle layer |
|
|
EMAGE:7983
view entry |
wild-type |
- |
ISH |
 |
|
|
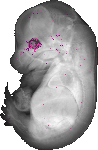
Ckmt1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.014 |
| Go to annotation details |
 |
 |
vagus X ganglion |
 |
rectum |
 |
vestibulocochlear VIII ganglion |
|
 |
glossopharyngeal IX ganglion |
 |
ventral grey horn |
 |
trigeminal V ganglion |
 |
nasal cavity olfactory epithelium |
|
|
EMAGE:10918
view entry |
wild-type |
- |
ISH |
 |
|
|
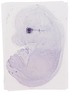
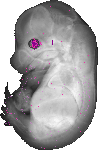
Cryba1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.013 |
|
EMAGE:8504
view entry |
wild-type |
- |
ISH |
 |
|
|
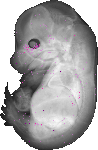
Dicer1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.012 |
|
EMAGE:10338
view entry |
wild-type |
- |
ISH |
 |
|
|
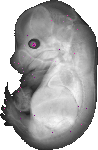
Crybb2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.012 |
|
EMAGE:12006
view entry |
wild-type |
- |
ISH |
 |
|
|
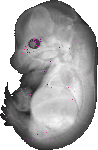
Rtp2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
|
EMAGE:14337
view entry |
wild-type |
- |
ISH |
 |
|
|
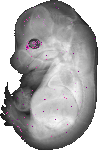
Foxn4 |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
|
EMAGE:13829
view entry |
wild-type |
- |
ISH |
 |
|
|
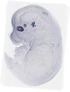
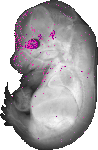
Rax |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
|
EMAGE:10998
view entry |
wild-type |
- |
ISH |
 |
|
|
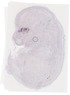
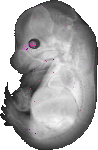
Atp6v1c2 |
 |
 |
 |
TS23 |
14.5 dpc |
0.011 |
|
EMAGE:21443
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
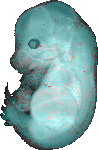 download query region as wlz file
download query region as wlz file