|
|
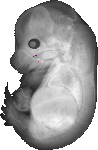
Pus7 |
 |
 |
 |
TS23 |
14.5 dpc |
1.000 |
- |
EMAGE:18163
view entry |
wild-type |
- |
ISH |
 |
|
|
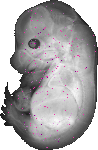
F2r |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
|
EMAGE:19217
view entry |
wild-type |
- |
ISH |
 |
|
|
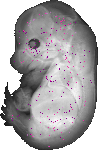
Wdr89 |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
- |
EMAGE:19842
view entry |
wild-type |
- |
ISH |
 |
|
|
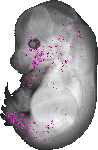
Kera |
 |
 |
 |
TS23 |
14.5 dpc |
0.007 |
| Go to annotation details |
 |
 |
extrinsic ocular muscle |
 |
olfactory cortex marginal layer |
 |
hand mesenchyme |
|
 |
upper lip |
 |
lower lip |
 |
nasal septum |
 |
palatal shelf |
 |
cranial muscle |
 |
foot mesenchyme |
 |
submandibular gland primordium |
|
|
EMAGE:12002
view entry |
wild-type |
- |
ISH |
 |
|
|
Sox7 |
 |
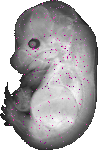 |
 |
TS23 |
14.5 dpc |
0.006 |
- |
EMAGE:14855
view entry |
wild-type |
- |
ISH |
 |
|
|
Iqgap1 |
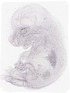 |
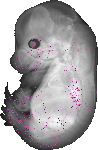 |
 |
TS23 |
14.5 dpc |
0.006 |
| Go to annotation details |
 |
 |
lung |
 |
metencephalon part of 4th ventricle choroid plexus |
 |
mandible |
|
 |
lateral ventricle choroid plexus |
 |
metanephros |
 |
hindgut |
 |
vibrissa |
 |
oral epithelium |
 |
orbito-sphenoid |
 |
nasal cavity epithelium |
 |
midgut |
|
|
EMAGE:8860
view entry |
wild-type |
- |
ISH |
 |
|
|
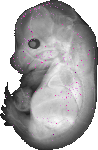
Avpi1 |
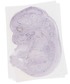 |
 |
 |
TS23 |
14.5 dpc |
0.006 |
| Go to annotation details |
 |
 |
mandible |
 |
basisphenoid bone |
 |
stomach |
|
 |
right lung |
 |
orbito-sphenoid |
 |
left lung |
 |
adrenal medulla |
 |
rib |
 |
trachea cartilaginous ring |
 |
clavicle |
 |
pharynx |
 |
maxilla |
|
|
EMAGE:11521
view entry |
wild-type |
- |
ISH |
 |
|
|
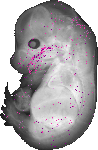
Pik3ip1 |
 |
 |
 |
TS23 |
14.5 dpc |
0.006 |
| Go to annotation details |
 |
 |
hindbrain meninges |
 |
midbrain meninges |
 |
oral epithelium |
|
 |
diencephalon meninges |
 |
telencephalon meninges |
|
|
EMAGE:19771
view entry |
wild-type |
- |
ISH |
 |
|
|
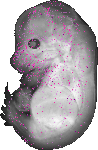
2510003E04Rik |
 |
 |
 |
TS23 |
14.5 dpc |
0.005 |
- |
EMAGE:14324
view entry |
wild-type |
- |
ISH |
 |
|
|
Itga10 |
 |
 |
 |
TS23 |
14.5 dpc |
0.005 |
| Go to annotation details |
 |
 |
fibula |
 |
tibia |
 |
axial skeleton cervical region |
|
 |
axial skeleton sacral region |
 |
otic capsule |
 |
pituitary gland |
 |
mandible |
 |
basioccipital bone |
 |
basisphenoid bone |
 |
Meckel's cartilage |
 |
tarsus |
 |
axial skeleton lumbar region |
 |
temporal bone petrous part |
 |
scapula |
 |
humerus |
 |
maxilla |
 |
viscerocranium |
 |
axial skeleton thoracic region |
 |
hip |
 |
nasal septum |
 |
orbito-sphenoid |
 |
femur |
 |
rib |
|
|
EMAGE:11783
view entry |
wild-type |
- |
ISH |
 |
 icon to keep this page displayed.)
icon to keep this page displayed.)
 icon to keep this page displayed.)
icon to keep this page displayed.)
 download query region as wlz file
download query region as wlz file